Table of Contents
- Overview
- Tagging ELN documents, folders, and notebooks
- Searching for Tagged content
- Tagging Rules
- Controlled vocabularies/ontologies
- Import Tags by Uploading ontology files in csv format
- Viewing Tag Metadata (for imported and external ontologies)
- Sharing ontologies
- BioPortal Ontologies
- Enforcing ontology tags usage in LabGroup
- Tags in Inventory
Tagging & Controlled Vocabularies
Overview. There are currently three possible sources for tags that can be used in both the ELN and the Inventory (RSINV) system. The types of tags you can use are configurable at the LabGroup level b…
- Overview
- Tagging ELN documents, folders, and notebooks
- Searching for Tagged content
- Tagging Rules
- Controlled vocabularies/ontologies
- Import Tags by Uploading ontology files in csv format
- Viewing Tag Metadata (for imported and external ontologies)
- Sharing ontologies
- BioPortal Ontologies
- Enforcing ontology tags usage in LabGroup
- Tags in Inventory
Overview
There are currently three possible sources for tags that can be used in both the ELN and the Inventory (RSINV) system. The types of tags you can use are configurable at the LabGroup level by the PI.
1) Tags can be "free text" - typed by users, but remembered and subsequently autocompleted by the system.
2) Tags can be autocompleted based on a list read from one or more managed documents created in RSpace using Create > from form > "RSpace Tags from Ontologies". These documents can be edited manually, or populated using CSV files imported using the My RSpace > Export-Import page.
3) Tags can also be selected from external industry standard ontologies such as https://bioportal.bioontology.org/ connected to RSpace via the RSpace API.
When adding tags, the system will make suggestions after you have typed a few characters:
PIs can decide to enforce the use of ontologies within their lab group, this will remove the option to enter tags as free text and enforce that all tags associated with documents MUST come from either ontology files shared with a Group, or, if allowed, from an external ontology. This is described in detail below.
Tagging ELN documents, folders, and notebooks
Tagging multiple records in Workspace listing
You can tag any document, folder or notebook that you can edit – just select the record(s) in Workspace listing, and click on 'Add/Remove Tags' action.
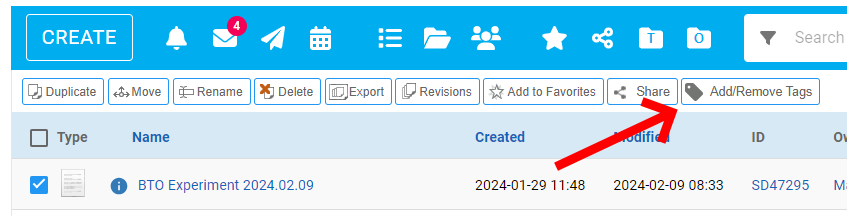
The tagging dialog allows for adding/removal of tags from all selected records. It starts up with showing tags that are applied to every selected record.
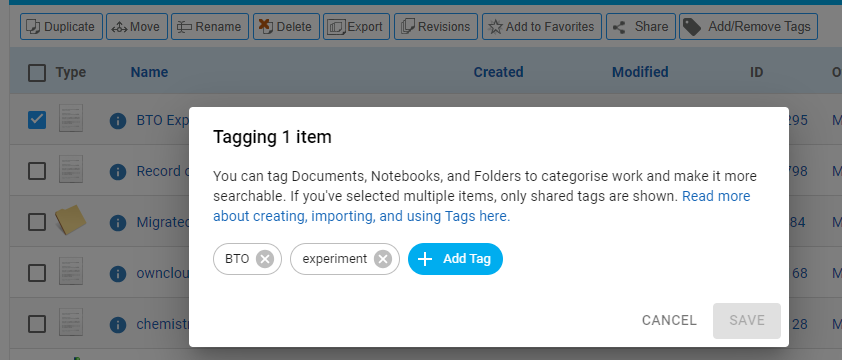
Deleting the tag will remove it from all selected records, adding new tag will add it to all selected records.
Tagging individual documents in Document View
Additionally, when viewing a document the current tags are displayed at the top-right-hand side of the document view, next to the name. To edit, just click on the 'edit' symbol and add them as a comma-separated list in the ‘Tags’ textbox .
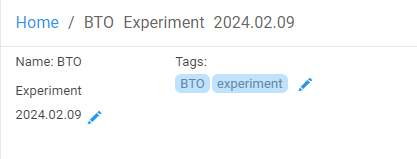
Clicking on 'edit tags' displays a text input which will be autopopulated with 'suggestions'. It will also display existing tags as tag 'pill's. Click on a pill to see information about that tag and or click on the 'x' in the pill to delete the tag.
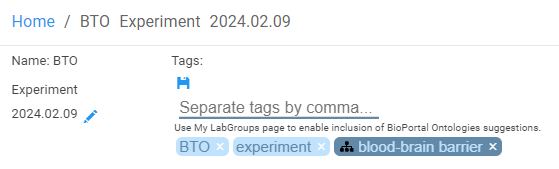
Searching for Tagged content
After clicking on 'filter' icon In Workspace search dialog, you can choose to search by tag(s):
When using tag search, RSpace autopopulates the possible search terms from existing tags:
Searching for tags is a very powerful way to organise and aggregate your research documents in multiple ways – for example by grant number, project, or publication, or simply as a collective name for a related or themed set of documents. To view only a selection of documents marked with a particular tag, choose ‘Tag’ in the Workspace search drop-down and enter the identifying term you have tagged a series of entries or documents with – the search returns a table of results which only shows items containing the particular tag. Tagging documents makes them easier to find and collects them in search results with similarly tagged related content.
Tagging Rules
Stop words and Tagging
There are various words that the search engine will not find. These are generally short prepositions such as 'of', 'and', 'this', 'that' etc. The search engine will not be able to find tags that are solely comprised of these stop words. It will, however, find tags that include these stop words but are not exclusively using stop words. Therefore it will not find a document tagged with the value: 'of'. It will find a document tagged with 'Ides of March'.
When creating a new tag RSpace suggests autopopulated values from existing tags AND from any ontology files in your workspace or shared with you (see below).
You may use free text instead. Simply click in the textbox for tags in order to see the suggestions.
Using key value pairs in Tags
Tags can be created as key=value - just enter the tag as 'key=value' (no surrounding quotes are required) using freetext or in an ontology file (see below). Searches for a tag with 'key=value1' will return only documents tagged with 'key=value1' and not documents tagged with 'key=value2' etc.
Forbidden characters in Tags
The following characters are forbidden in tags and will be rejected if entered as free text or from a tag suggestion term: '<', '>', '\'. A forward slash '/' is allowed from a suggestion but will be forbidden if entered as free text.
Controlled vocabularies/ontologies
Autogenerated tags ontology
Whenever you save (or delete) a tag, RSpace creates/updates a document inside an 'Ontologies' folder in your workspace with an icon representing its purpose:
This file is an example of a controlled vocabulary/ontology document. Do not edit this file by hand as it will be overwritten on any future saving of your tags. The file does not exist until you save/delete a tag. You may want to share or export this file, allowing other users access to the controlled vocabulary which you have created as document tags.
Creating vocabulary/ontology documents
You may create ontology files for the purpose of ensuring an agreed set of terms are used for tags. Although referred to as ontologies/controlled vocabularies throughout this documentation, these files are really just a controlled vocabulary as they have no concept of nested terms or hierarchies. However they do contain the ability to create key=value pairs which could be useful for e.g. namespacing.
To create an ontology file:
Click create -> From Form -> RSpace Tags from Ontologies
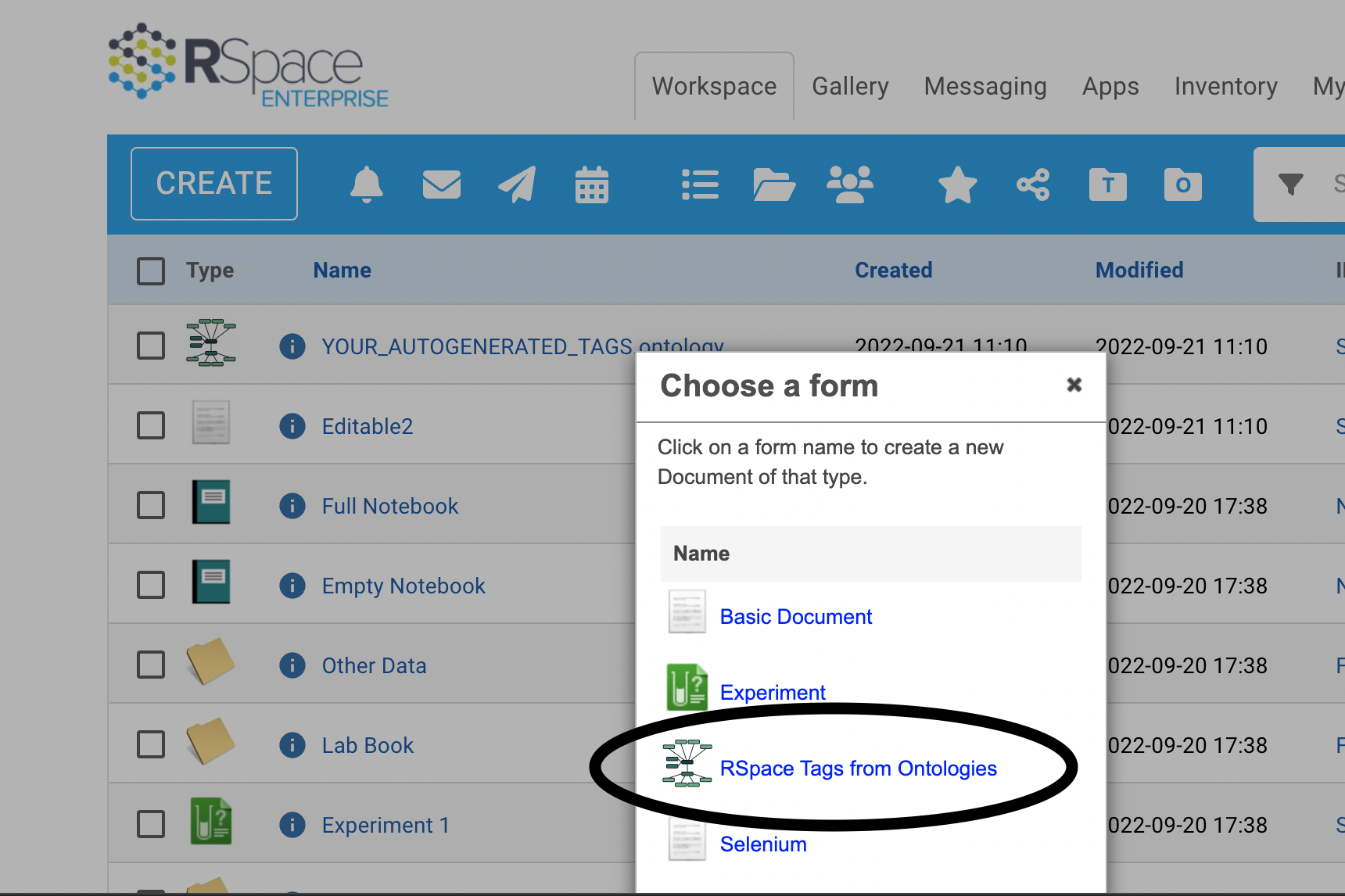
The generated ontology file contains 20 fields, each called 'Ontologies for Tag creation'.
This file is just a normal RSpace file but it will be used to generate tag 'suggestions' following some simple rules as follows:
Ontology terms should be comma separated. There can be one key per line of text, separated by an '=' from the values it matches.
For example, I create a controlled vocabulary to describe experiments so I edit 'Ontologies for Tag creation' and enter : "started,finished,phase1,phase2". Whenever I chose to tag a document, these 4 values will appear as 4 separate suggested terms for the new tag. If I wished to namespace this, then I would enter: "experiment_stage=started,finished,phase1,phase2". Whenever I chose to tag a document I will now have 4 separate suggested key=value pairs for the tag. These will be 'experiment_stage=started', 'experiment_stage=finished' etc.
There can only be one key per line of text, therefore in order to create further key=value pairs you must enter values on a seperate line in the file:
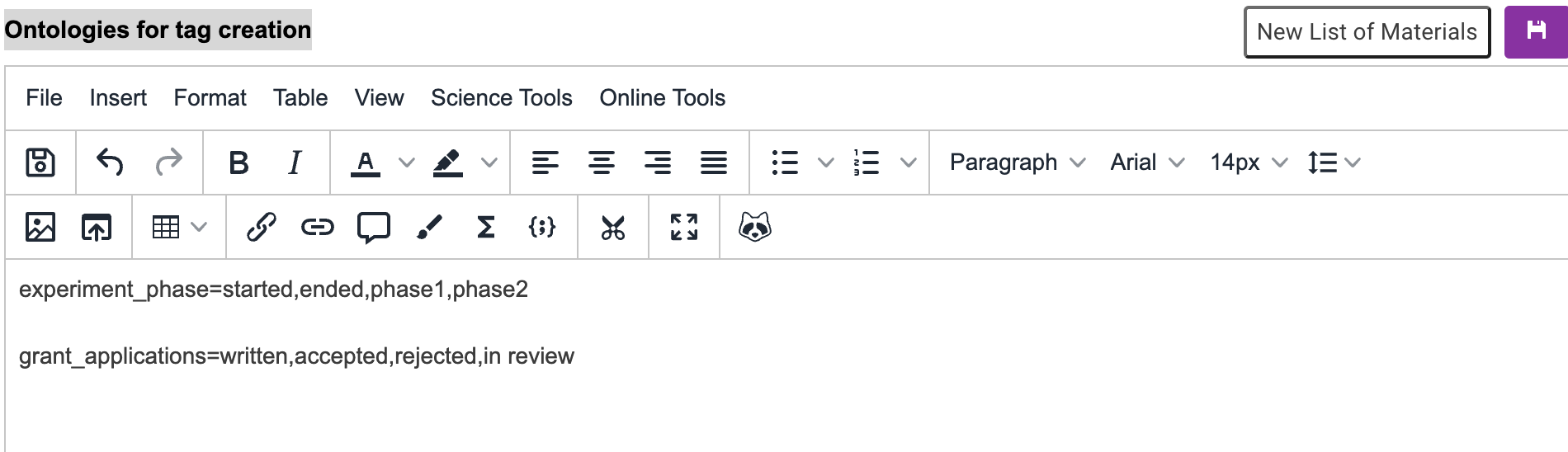
After saving the ontology document, whenever I create a new tag for any document I will see the following suggestions:
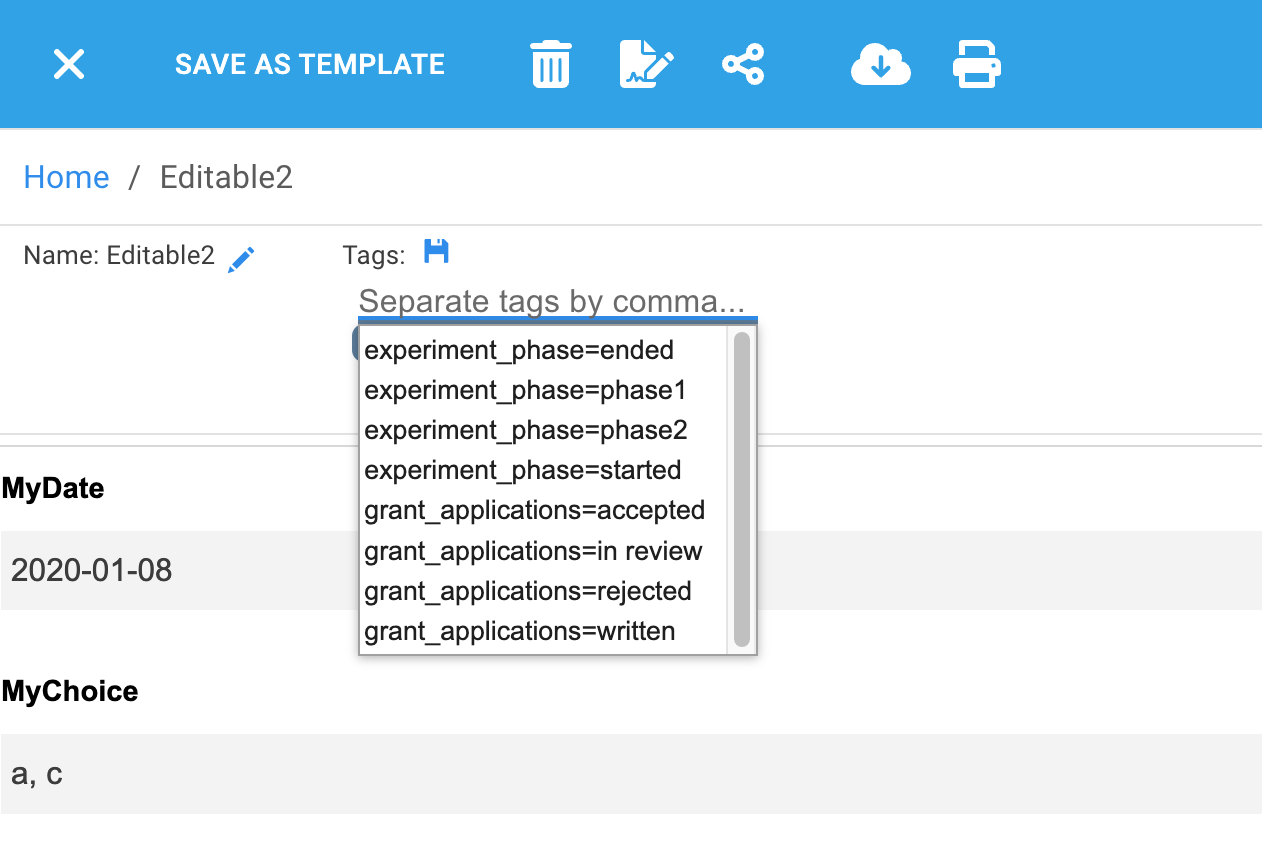
Listing visible ontology files
It can be useful to see all ontology files you own or are shared with you. There is an 'ontologies' view in the workspace:
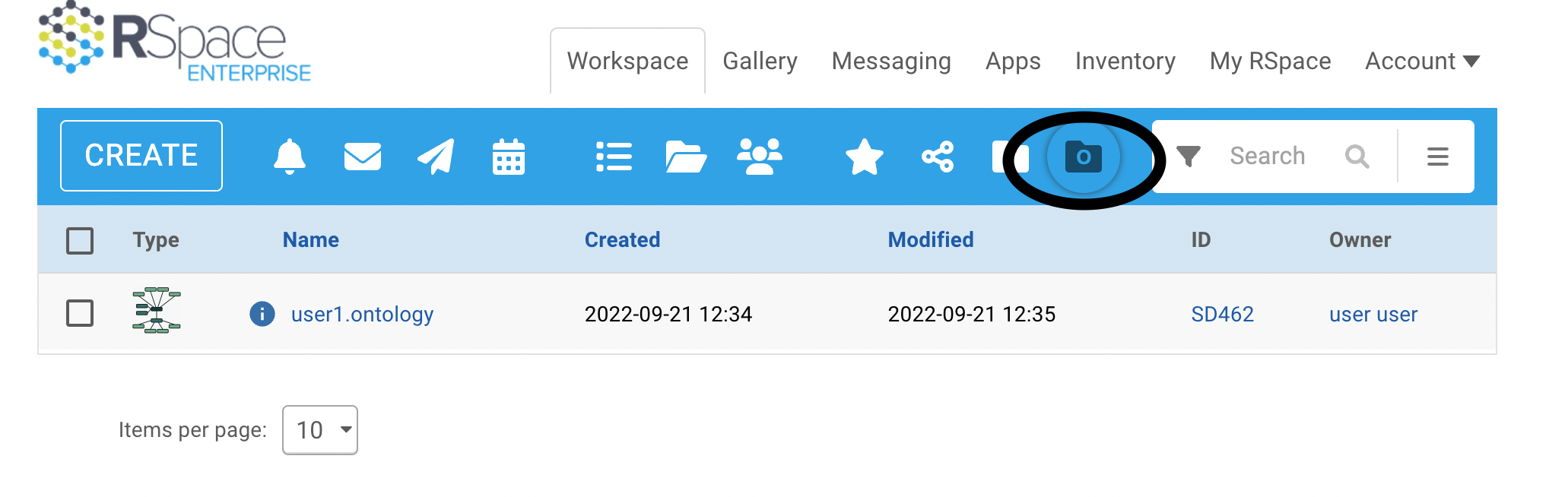
This shows your ontology files and ontology files shared with you. Note that if ontologies are enforced, only those files which have been shared with a Group will be making any contribution to the controlled vocabulary you can use to create new tags. Click on the info, 'i', button for any RSpace document to show you whether it has been shared with a Group.
Import Tags by Uploading ontology files in csv format
If an external ontology file is available in CSV format, it can be uploaded to RSpace. For example https://bioportal.bioontology.org/ allows download of ontologies in the CSV format. This method can also be used to quickly and efficiently used to create a long list of tags based on some pre-existing list of terms the user has access to but the list needs to be formatted as a CSV file with one tag / term / text value per row.
Example: The Helmholtz Metadata Collaboration provides a standardized Electron Microscopy Glossary (EMG) with harmonized terminology for electron microscopy research. The EMG vocabulary is available as a CSV file at https://codebase.helmholtz.cloud/em_glossary/em_glossary_owl/-/blob/main/emg.csv and includes terms like "acquisition time", "dwell time", "scattering angle", and "working distance" that can be imported as controlled vocabulary tags for electron microscopy experiments.
To import a CSV, go to the export-import page under the My RSpace tab and choose a file using the dialog under 'Import an ontology file - csv format'.
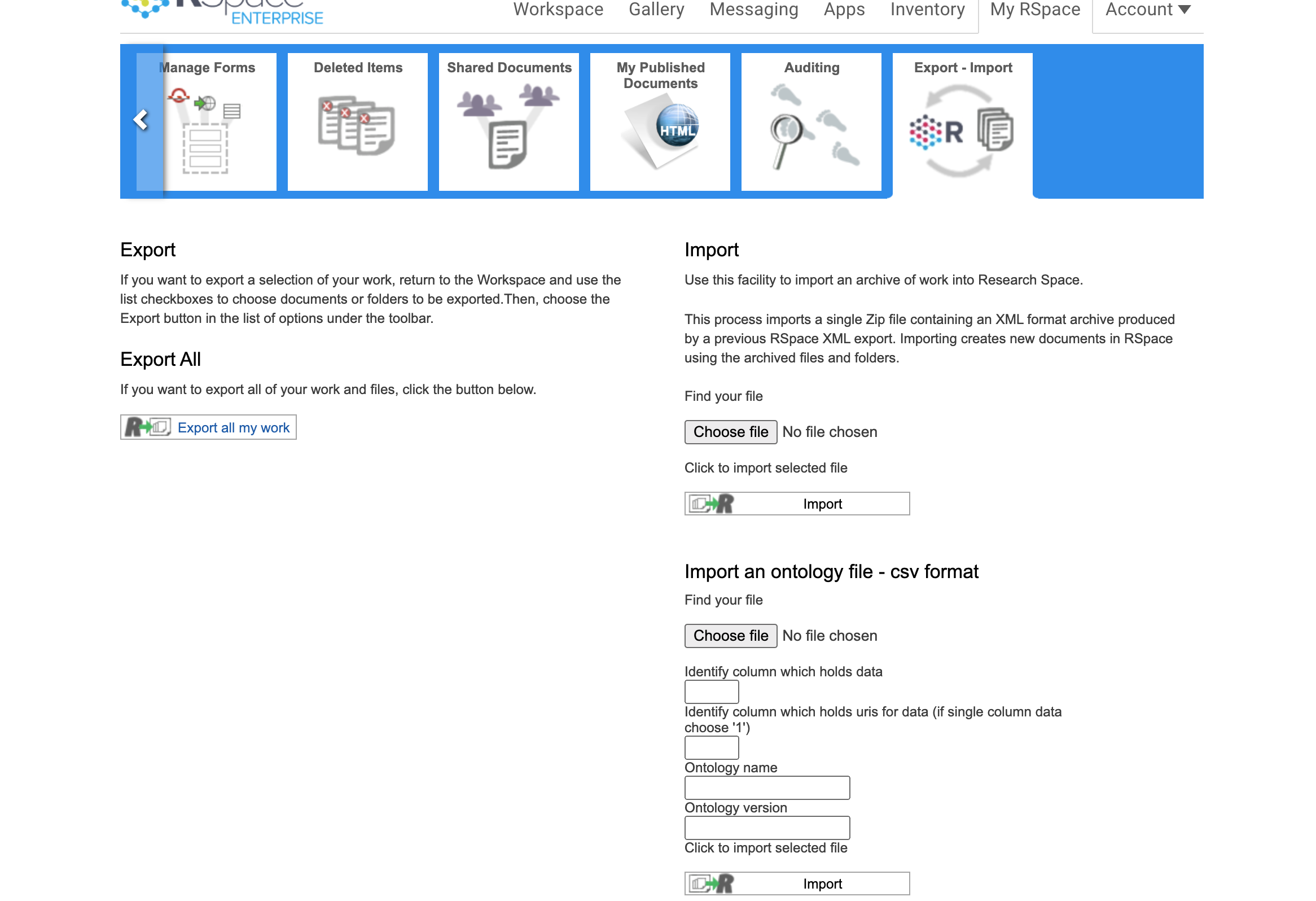
All inputs on the form are mandatory.
You must choose a single column that will be used as the list of terms for the controlled vocabulary. You must also choose a column which contains URIs to identify the source for ontology term. For example, having downloaded the 'BRENDA Tissue and Enzyme Source Ontology' from https://bioportal.bioontology.org/ontologies/BTO in CSV form, the user can decide to use the 'preferred label' column, which is column 2 in the CSV file. I enter 2 under 'Identify column which holds data'. URIs are contained in column '1' so I enter 1 under 'Identify column which holds URIs for data'). You can also add the official ontology name and official ontology version which should be available on the third party site the ontolology came from. The CSV file should contain only the data you want to import, so if the first row includes headings, you will need to delete that row of the CSV.
Note that you may optionally import an Ontology which consists of only a single column of values. In that case, enter '1' for both fields. This is useful when working with a long list of terms that you don't what to manually type into an RSpace ontology document and / or if there is no URI or other metadata about the origin of the terms in the list.
After a succesful upload, RSpace will open the workspace, showing the new file. The ontology terms will all be on one 'line', with up to 10,000 terms per 'Ontologies for Tag creation' field in the document. There are 20 fields in the document, which means an uploaded CSV file can contain at most, 200000 ontology terms.
If your CSV included both terms AND URI metadata for each term, then the imported document will look something like this in RSpace:
Note that RSpace ontology files created using this import method are automatically signed on import in order to protect the integrity of third party controlled vocabularies. If you need to make edits to an imported ontology you can do so, but you will need to select the ontology file and click "duplicate" to make a new unsigned copy. You can edit the new copy, but not the original.
Viewing Tag Metadata (for imported and external ontologies)
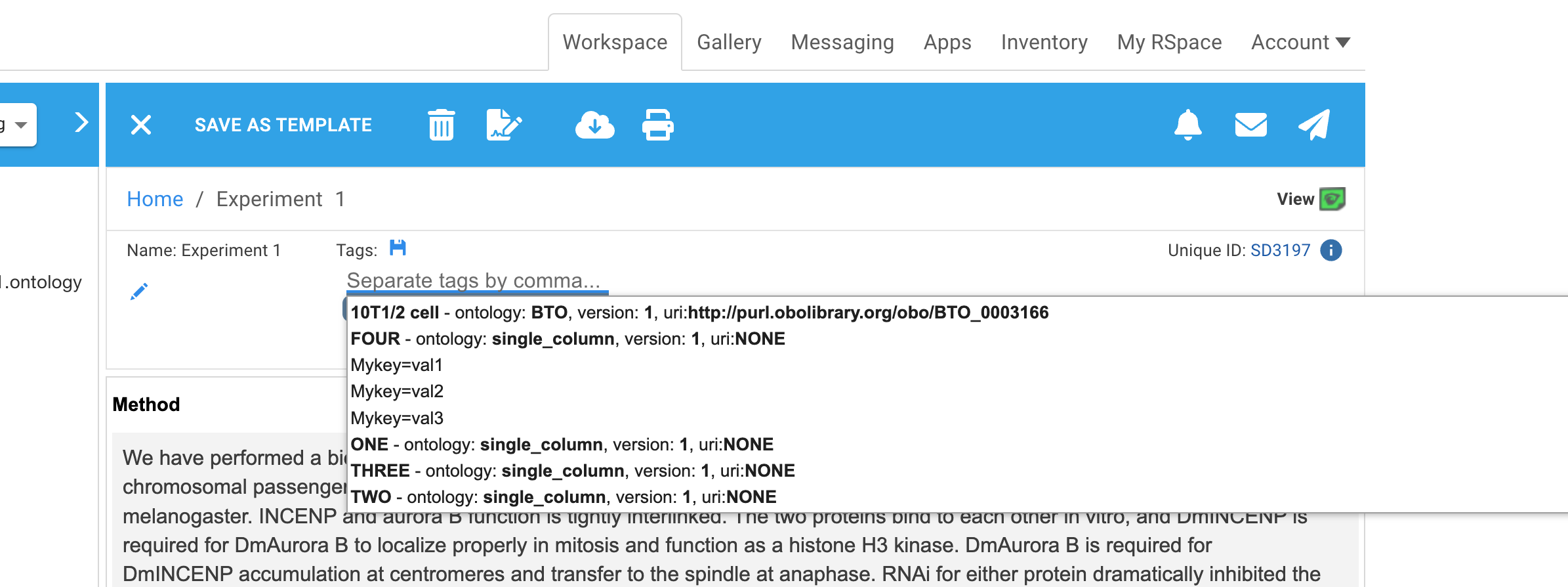
Ontology terms from imported CSV files and external ontologies can show associated metadata so that their source can be determined. For imported CSV files this includes the name and version you gave the upload as well as the associated URI for the term. Metadata is displayed in tag suggestions:
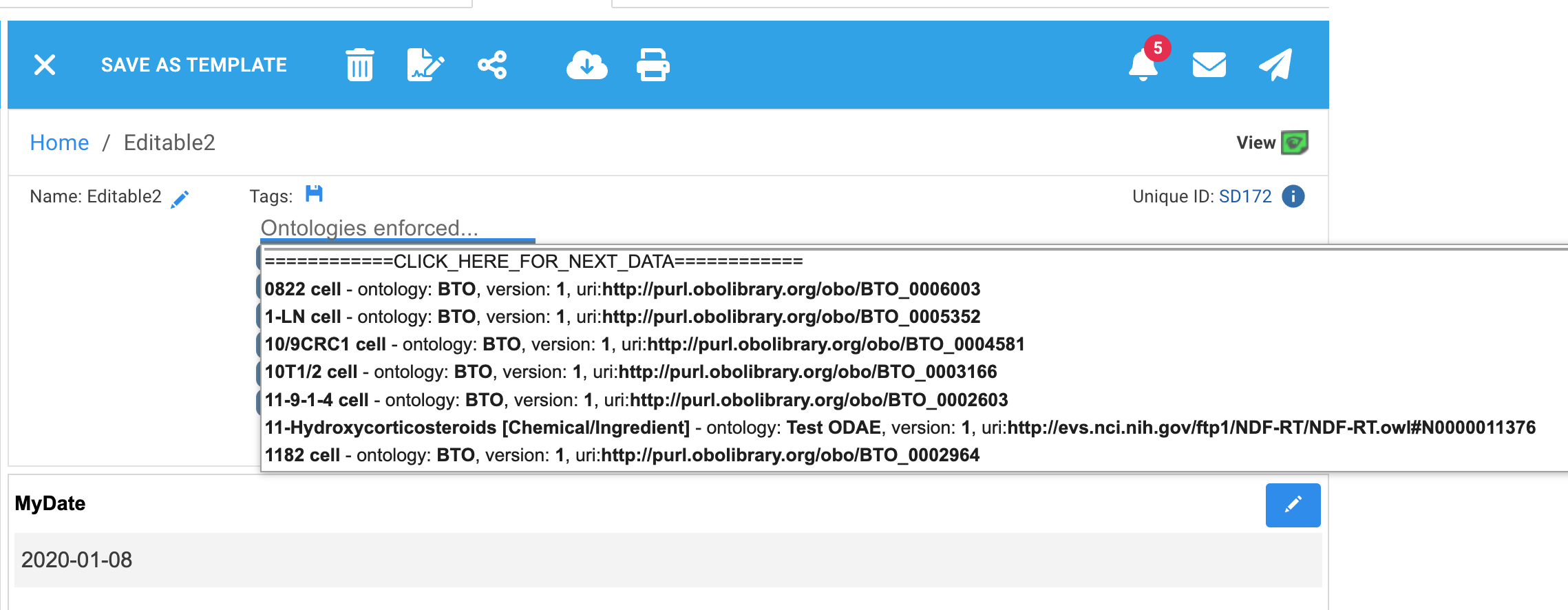
If the ontology file had only a single column of data, URIs will be displayed as 'NONE'
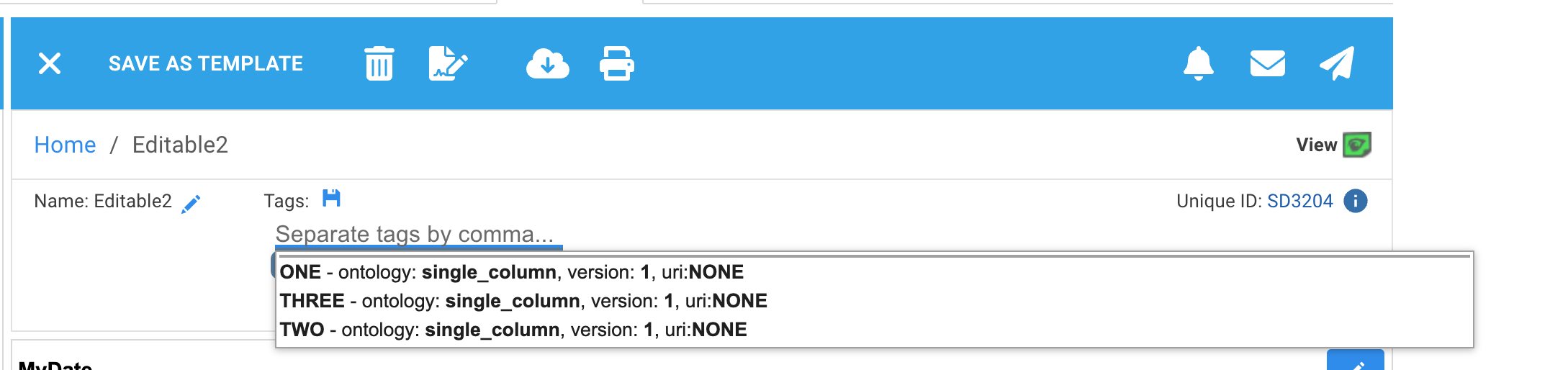
Ontology terms from freetext or from your own manually created ontology documents does not have any meta data displayed in suggestions:
Metadata is also displayed when you click on a tag 'pill'.
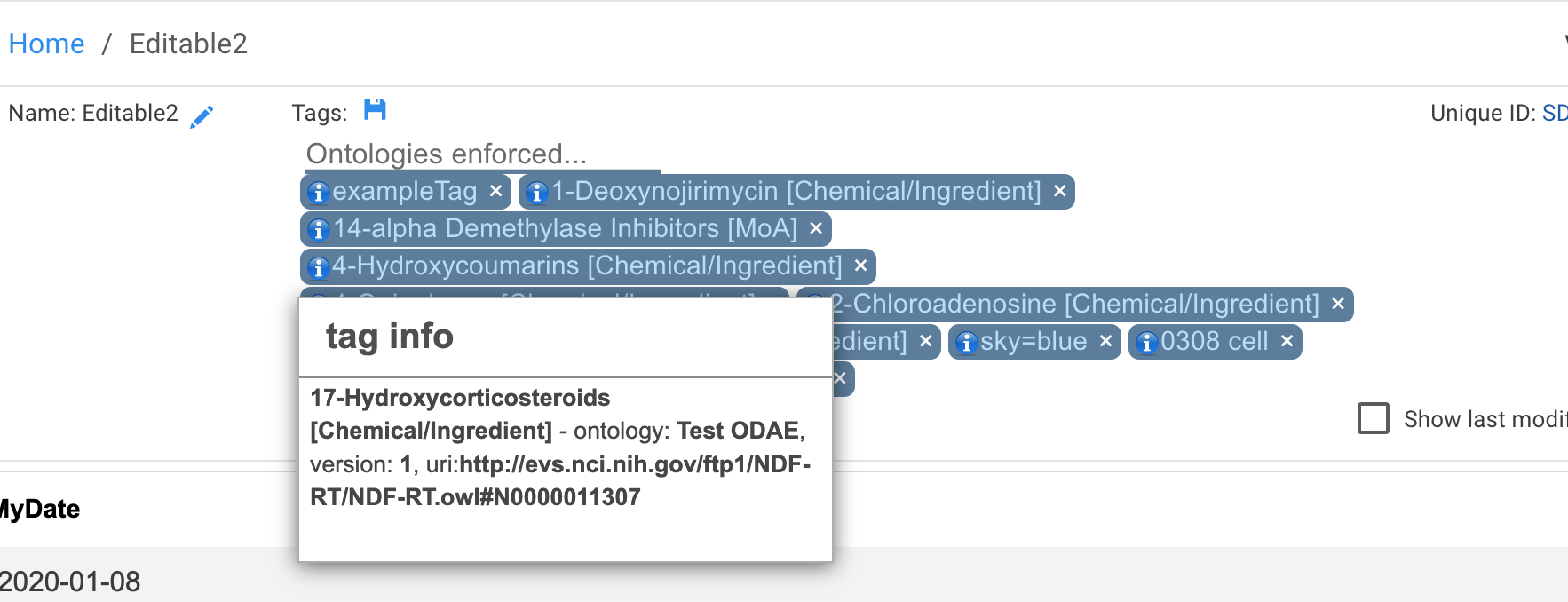
Tag info popups will be dismissed by subsequent clicks anywhere on the screen. Opening the tag info popup above would copy the tag URI - "http://evs.nci.nih.gov/ftp1/NDF-RT/NDF-RT.owl#N0000011307" to the clipboard. You can use this to search for the URI in your browser.
Metadata is also saved to your autogenerated ontology file whenever you add a tag to a document. It is also saved to the tagged document itself. Therefore metadata associated with a tag saved to a document will not be lost should the ontology file that tag came from be deleted/unshared etc.
Sharing ontologies
Ontology files can be shared as with any RSpace document. Recipients will be able to use them to create tags based on those ontology documents. You can share RSpace ontologies with LabGroups, Collaboration Groups and Project groups, and the owner can choose to share with either read or edit access. This allows for tremendous flexibility and helps groups of users categorize work in similar and consistent ways. This also allows for specialization in the categoriztion of work for different segments of your institution. For example, chemistry labs can use ontology files focussed on categorizing types of chemical reactions and instructors can catgorize work based on which class the work is used in. By sharing ontology files appropriately, users will not be overwhelemed by lists of tags that are not relevent to their own work.
You may export an ontology file as an RSpace archive. Any recipient can then re-import it (i.e. as an RSpace archive, not as a CSV file).
BioPortal Ontologies
You can use data from the BioPortal Ontologies site as a source of tag suggestions.
Whetzel PL, Noy NF, Shah NH, Alexander PR, Nyulas C, Tudorache T, Musen MA. BioPortal: enhanced functionality via new Web services from the National Center for Biomedical Ontology to access and use ontologies in software applications. Nucleic Acids Res. 2011 Jul;39(Web Server issue):W541-5. Epub 2011 Jun 14.
- Group PIs can chose to allow "BioPortal Ontologies" on the MyLabGroups page. By default this is not allowed.
- Group members, including PIs, can only use BioPortal Ontologies if ALL groups to which they belong have allowed use.
- Once allowed, data from BioPortal Ontologies will be shown among your own Ontologies as tag suggestions.
- When creating new tags, you must enter a search term of > 2 characters long before the BioPortal Ontologies data will be queried and shown among your tag suggestions.
- Tip - use a discriminative term when searching, there can be many results returned otherwise.
Enforcing ontology tags usage in LabGroup
In order to guarantee all group members use a shared vocabulary for tagging, a Lab Group PI has the option to enforce ontologies on the My LabGroups page:
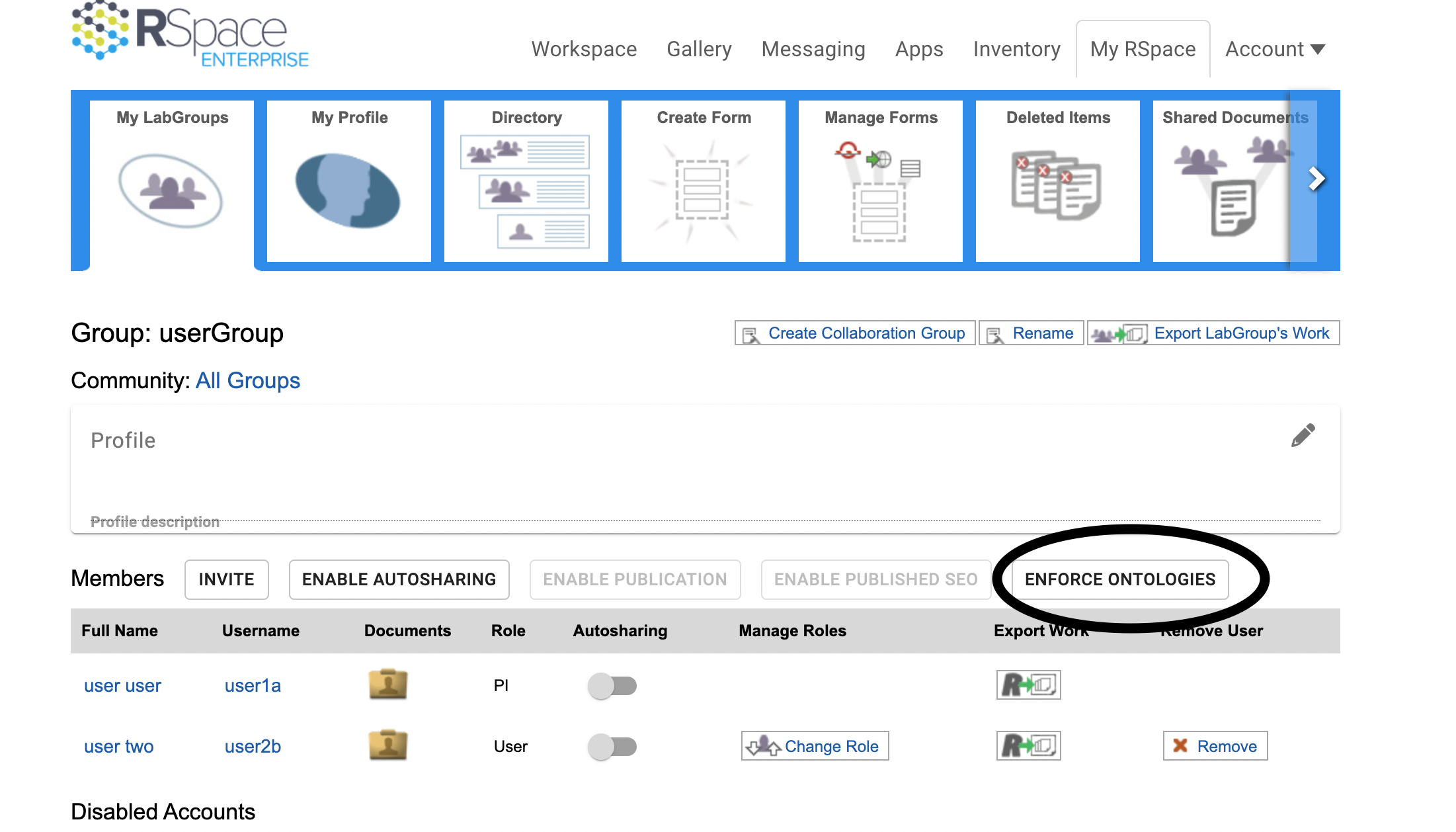
If ontologies are enforced the following applies to all members of the Lab Group/ Collaboration Group (including the PI):
New Tags cannot be entered as free text.
New Tag suggestions will not be autopopulated from existing tag values.
New Tag suggestions will only be autopopulated from ontology files which have been shared with a Group.
As a user, if any group I am a member of has 'enforce ontologies' turned on, then these rules take affect in my workspace. This includes collaboration groups.
Some examples:
- I am group PI and I turn on Enforce Ontologies. No ontology files have been shared with any group I belong to. I will not be able to create any new tags until an ontology file is shared with a group I belong to.
- I am group PI and I belong to a collaboration group. Another PI in the collaboration group turns on 'enforce ontologies' for the collaboration group. No ontology files have been shared with any group I belong to. I will not be able to create any new tags until an ontology file is shared with a group I belong to.
- I am a member of a group with enforced ontologies and I wish to use my own ontology file to create tags. I share it with the PI. I will not be able to use the ontology file to create tags and neither will the PI. I must share the file with a Group before I, or any member of the group, may use the file to create tags.
Using tags within LabGroup that enforces ontologies
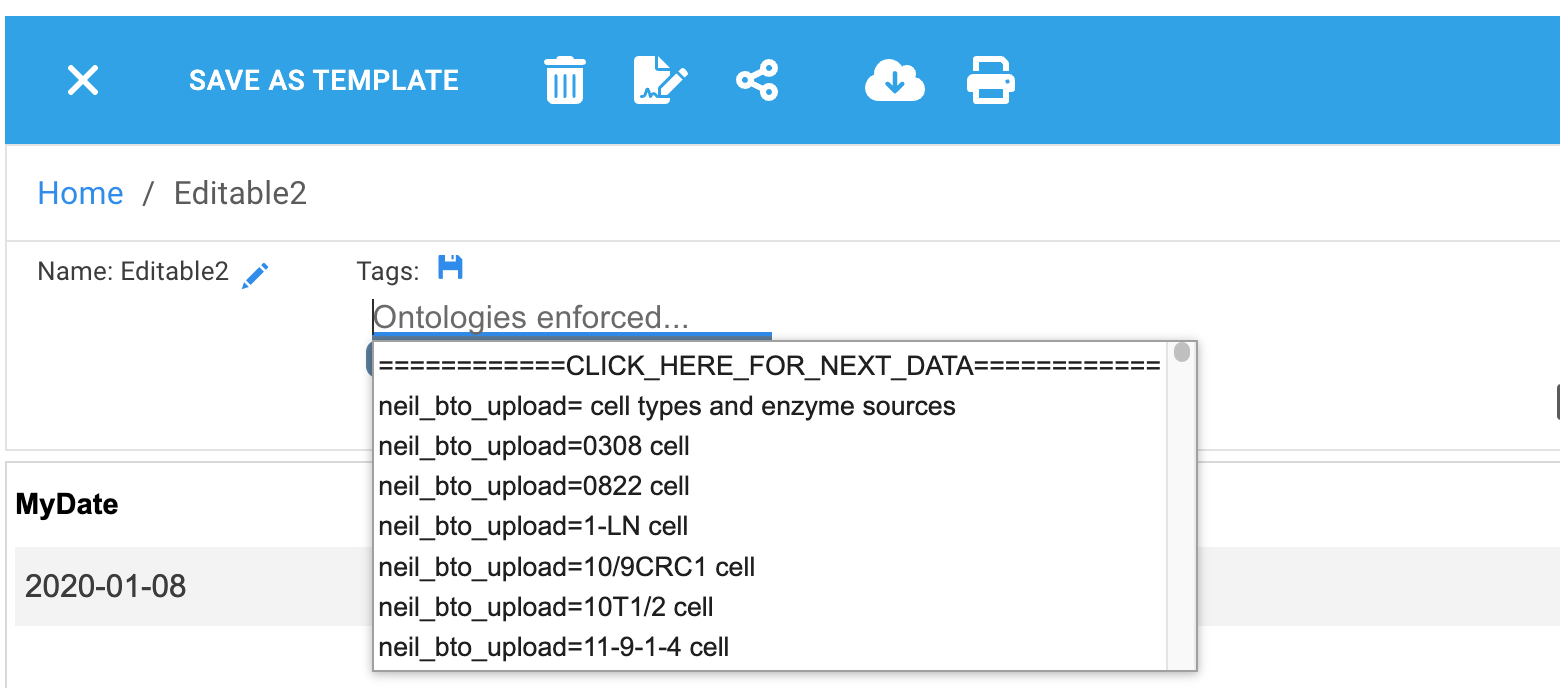
When ontologies are enforced, RSpace will only allow tag values from ontology files shared with a group you belong to. If you type in the tag text box this is used to filter the autopopulated suggestions. Suggestions are in alphabetical order. RSpace will only display up to 1000 suggestions, when there are more possible values than 1000, RSpace will display an initial value of
'============CLICK_HERE_FOR_NEXT_DATA============'.
Click on this value in the suggestions dropdown to load the next 1000 suggestions. When there is no more suggested data, RSpace will display:
'================BACK_TO_START================'
Click on this value in the suggestions dropdown to cycle back to the original suggestions.
When there are too many possible values, RSpace requires you to narrow them down by entering some text and will display: 'Too many results, please enter a specific search term'.
Tags in Inventory
Tags in Inventory support all of the functionality described above: imported controlled vocabularies, metadata, and the BioPortal. The interface is slightly different, but behaves much the same.
Adding tags
When creating a new item (a container, a sample, etc) or when editing an existing record you can tap "Add Tag" in the Tags section under Details, and the tag selection box will appear.
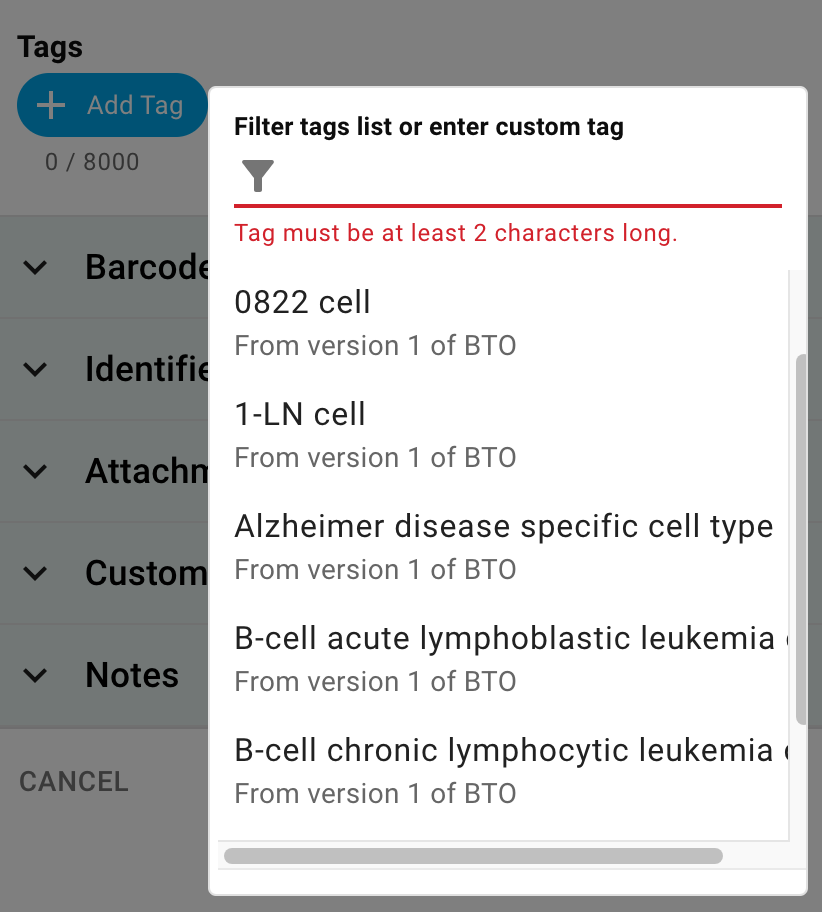
Here, you will see the same set of suggestions as when tagging documents in the ELN. Tags imported from controlled vocabularies -- be they imported CSV files or the BioPortal -- will have metadata about their version and ontological name. Once saved, you can view this metadata by tapping on the little blue icon.
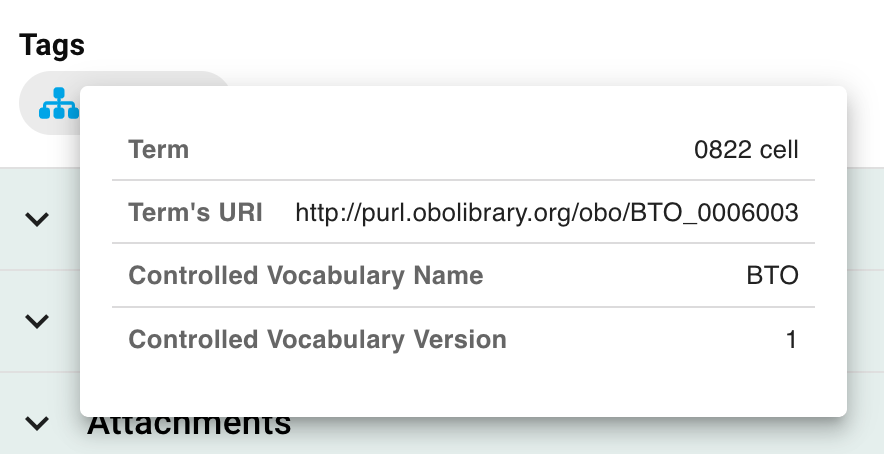
You are also free to enter any tag you like provided it is at least 2 characters in length, and does not contain any of the forbidden characters described above.
Searching for Tags
There are two ways to search for Inventory items that have a particular tag.
The first is you can search for it by typing into the "Tags" search parameter using the same menu as when adding tags.
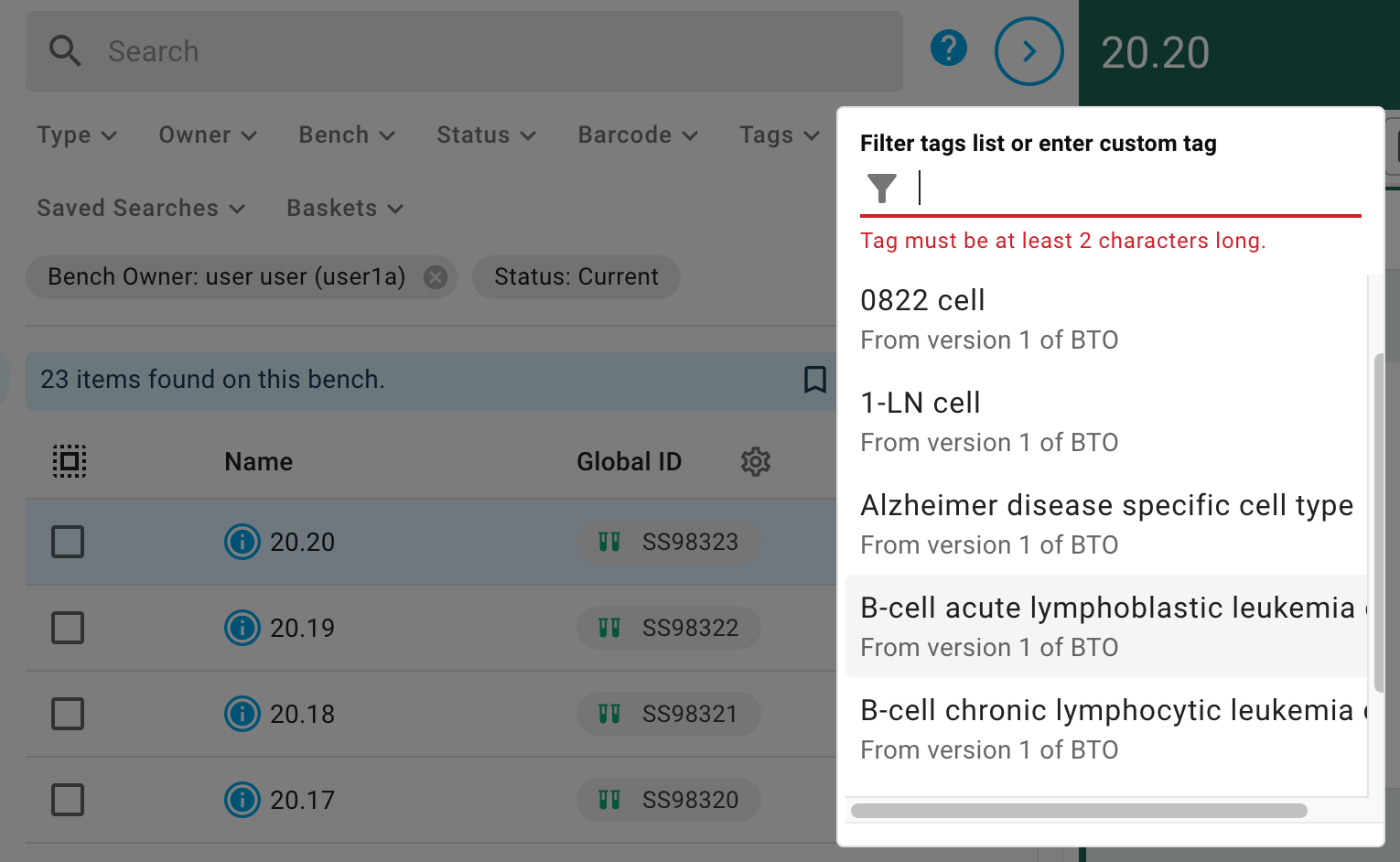
The second is to tap a Tag wherever they appear on the page, which will perform a search as if you had chosen the tag in the search menu.
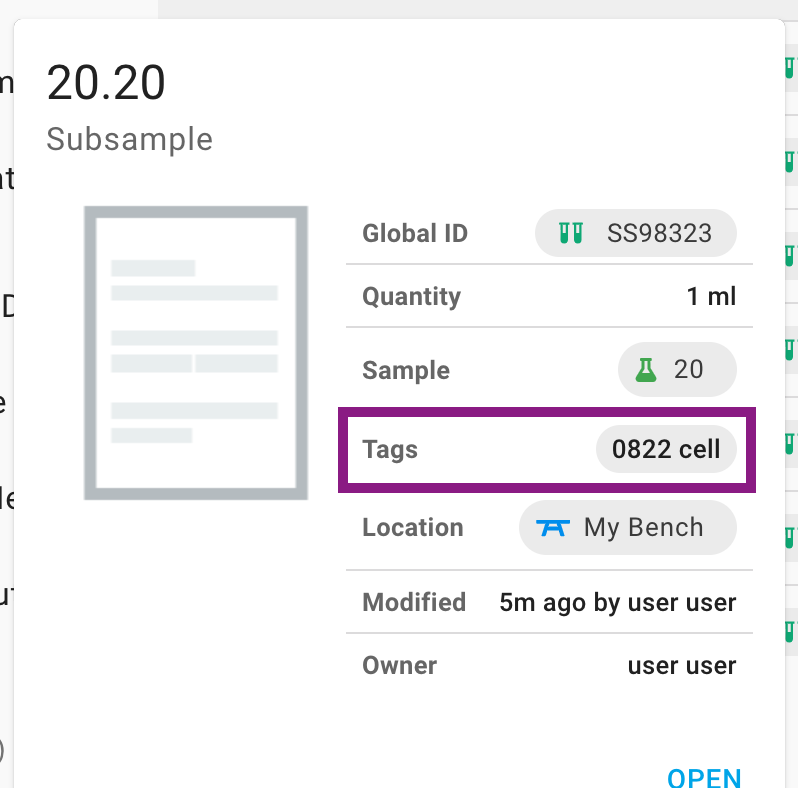
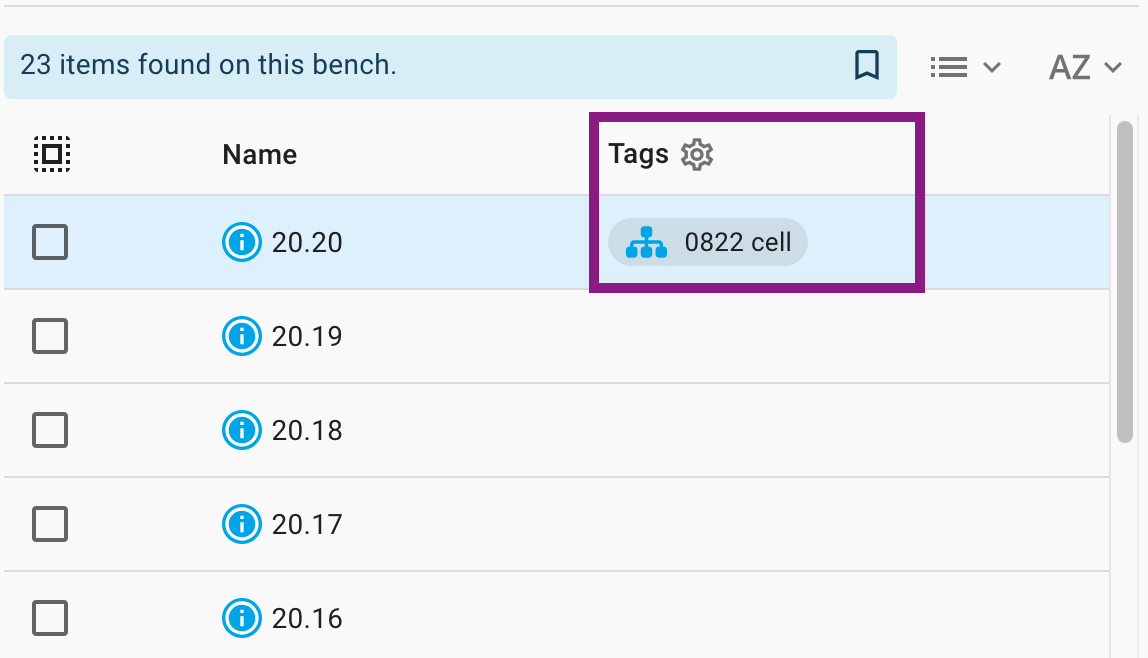
Do note that performing a seach for tags will trigger a Lucene query which allows for the tag search to be combined with a search for other parameters. For example, you could search for all the records that have a particular tag and are owned by you.
How did we do?
Revisions and Deletions
Folders
